| Dataset wrap |
|---|
| Rowcaps |
|---|
| Rows 1-2: | Show 2 records for TSPARMCD = "GLPTYP", using TSSEQ to indicate multiple records, since both GLP types apply for this example study. | | Row 3: | Shows that this study was conducted as a GLP study. | | Rows 4-5: | Show the study start date and study title. | | Rows 6-7: | Show the version of SEND Implementation Guide and version of Controlled Terminology used in this study. | | Row 8: | Shows the applicant's organization. | | Row 9: | Shows that the applicant's study reference ID is not applicable. | | Rows 10-13: | Show that TSGRPID has been used to link records (name, location, country) related to the test facility (TSGRPID = 1). The study director is associated with the test facility. | | Rows 14-16: | Show that TSGRPID (TSGRPID=2) has been used to link the information on the testing guideline followed on this study (TSTGDNAM, TSTGDORG, TSTGDVER). | | Shows the study type for this study. | | Shows that this study includes a Mammalian Cell Micronucleus Assay. | | Rows 19-20: | Show that the species is human and the cell line is TK6 lymphoblastoid in this study. |
|
| Dataset2 |
|---|
| Expand |
|---|
| title | ts.xpt trial summary (study level parms) |
|---|
| | Expand |
|---|
| 1 | TS | 1 | SSTYP | Study Type | REPEAT DOSE TOXICITY | 2 | TS | 1 | SPECIES | Species | RAT | 3 | TS | 1 | STRAIN | Strain/Substrain | FISCHER 344 | 4 | TS | 1 | SPLRNAM | Test Subject Supplier | HARLAN | 5 | TS | 1 | SDESIGN | Study Design | CROSSOVER | 6 | TS | 1 | ROUTE | Route of Administration | ORAL | 7 | | Row | STUDYID | DOMAIN | TSSEQ | TSGRPID | TSPARMCD | TSPARM | TSVAL | TSVALNF |
|---|
| 1 | 123 |
| TS | 1 |
| GLPTYP | Good Laboratory Practice Type | FDA |
1 | EXPSTDTC | Experimental Start Date | 2008-01-01 | | 2 |
| GLPTYP | Good Laboratory Practice Type | OECD |
| | 3 | 123 |
EXPENDTC | Experimental End Date | 2008-03-07 | 10 | TS | 1 | TRMSAC | Time to Terminal Sacrifice | P42D | 11 | TS | 1 |
| STSTDTC | Study Start Date |
12 | TS | 1 | DOSDUR | Dosing Duration | P42D | 13 | Example a Crossover study in the Rat with 3 dose levels and 3 dosing periodsthe in vitro genotoxicity potential using the in vitro Neutral Red Uptake assay |
| | 6 | 123 |
| TS | 1 |
| SNDIGVER | SEND Implementation Guide Version |
| TOBACCO IMPLEMENTATION GUIDE VERSION |
| 123 | TS | 1 |
| SNDCTVER | SEND Controlled Terminology Version | SEND Terminology 2021- |
16 | TS | 1 | STCAT | Study Category | TOX | 17 | SSPONSORSponsor Organization Sponsor 18SPREFIDSponsor's | Study Reference ID |
| NOT APPLICABLE | | 10 |
| 123 | TS | 1 | 1 | TSTFNAM | Test Facility Name | Example |
| 123 | TS | 1 | 1 | TSTFLOC | Test Facility Location | 10 Somewhere Street, Montgomery, AL 10000 |
| | 12 |
| 123 | TS | 1 | 1 | TFCNTRY | Test Facility Country | USA |
| | 13 |
AGETXT | Age Text | 6-8 | 23| Study Director | Dr. R. Smith |
| | 14 | 123 | TS | 1 |
AGEU | Age Unit | WEEKS | 24| 2 | TSTGDNAM | Testing Guideline Name | GUIDELINE FOR THE TESTING OF CHEMICALS No. 487 |
| | 15 | 123 | TS | 1 | 2 |
STDIR | Study Director | Dr. R. Smith | | Testing Guideline Organization | OECD |
| | 16 | 123 |
TRT | Investigational Therapy or Treatment | Drug A | | 2 | TSTGDVER | Testing Guideline Version | 29-July-2016 |
| | 17 | 123 |
TRTV | Treatment Vehicle | Saline | 27| Study Type | GENOTOXICITY IN VITRO |
| | 18 | 123 | TS | 1 |
2 | TRTVDESC | Treatment Vehicle Structured Description | 100.46 %(w/v) ISOTONIC SODIUM CHLORIDE SOLUTION {UNII VR5Y7PDT5W} |
| GNTXAID | Genetic Toxicology Assay Identifier | MNvit |
| | 19 | 123 |
TRTCAS | Primary Treatment CAS Registry Number | TEMPORARILY UNAVAILABLE |
|
| Expand |
|---|
|
| Row | STUDYID | ASSAYID | DOMAIN | ARMCD | ARM | TAETORD | ETCD | ELEMENT |
|---|
1 | TA | ARM01-Prod1 | Cigarette Smoke Condensate, 10 ug/mL | 1 | TREATMT | Treatment | 9 | 9 | SLS-110 | 1 | | Expand |
|---|
| title | te.xpt (trial elements) |
|---|
|
| Row | STUDYID | ASSAYID | DOMAIN | ETCD | ELEMENT |
|---|
TE | TREATMT | Treatment | | Expand |
|---|
|
| Expand |
|---|
|
Red font (CT is coming!, POSITIVE CONTROL, VEHICLE CONTROL, etc.)
ARMCD. TREATMENT (element). SETCD
ARM01 TREATMT. CS-01. (Cigarette Smoke 10 ug/mL)
ARM01
TA.XPT
TRIAL ARMS
| Row | STUDYID | ASSAYID | DOMAIN | ARMCD | ARM | TAETORD | ETCD | ELEMENT |
|---|
1 | TA | ARM01-Prod1 | Cigarette Smoke Condensate, 10 ug/mL | 1 | TREATMT | Treatment | 2 | TA | ARM02-Prod1 | Cigarette Smoke Condensate, 50 ug/mL | 1 | TREATMT | Treatment | 3 | TA | ARM03-Prod1 | Cigarette Smoke Condensate, 75 ug/mL | 1 | TREATMT | Treatment | 4 | TA | 4 | 1 | 5 | TA | 5 | 1 | 6 | TA | 6 | 1 | 7 | 7 | 1 | 8 | 8 | 1 | 9 | 9 | SLS-110 | 1 | 10 | ARM10 | SLS-200 | 1 | TE.XPT
TRIAL ELEMENTS
| Row | STUDYID | ASSAYID | DOMAIN | ETCD | ELEMENT |
|---|
TE | TREATMT | Treatment | tx.spt
TRIAL SET
example: Each plate has a separate product
Product, Smoking Regime (e.g. Traditional combustible, ENDS), Smoke Fraction, etc.
https://www.itis.gov/servlet/SingleRpt/SingleRpt?search_topic=TSN&search_value=969610#null
| Row | STUDYID | ASSAYID | DOMAIN | SETCD | SET | TXSEQ | TXPARMCD | TXPARM | TXVAL |
|---|
1 | CS-10 | Cigarette Smoke Condensate, 10 ug/mL | METACT | Metabolic Activation | +S9 | 2 | CS-10 | Cigarette Smoke Condensate, 10 ug/mL | METACT | Metabolic Activation | -S9 | TX | CS-10 | Cigarette Smoke Condensate, 10 ug/mL | ARMCD | Arm Code | CS-10 | TX | CS-10 | Cigarette Smoke Condensate, 10 ug/mL | SPECIES | Species | Salmonella enterica | TX | CS-10 | Cigarette Smoke Condensate, 10 ug/mL | STRAIN | Strain/Substrain | Salmonella enterica enterica | TX | CS-10 | Cigarette Smoke Condensate, 10 ug/mL | PRODUCT | Product | product A | TX | CS-10 | Cigarette Smoke Condensate, 10 ug/mL | REGIME | Smoking Regime | Traditional combustible | TX | CS-20 | Cigarette Smoke Condensate, 50 ug/mL | ARMCD | Arm Code | CS-20 | TX | CS-20 | Cigarette Smoke Condensate, 50 ug/mL | PRODUCT | Product | product A | ... | TX | LSL-200 | sodium lauryl sulphate (SLS) positive control, 200 ug/mL | TRTDOS | Dose Level | 100 | TX | LSL-200 | sodium lauryl sulphate (SLS) positive control, 200 ug/mL | TCNTRL | Control Type | POSITIVE CONTROL | TX | LSL-200 | sodium lauryl sulphate (SLS) positive control, 200 ug/mL | CNTLAG | Control Agent | SODIUM LAURYL SULPHATE | TX | LSL-200 | sodium lauryl sulphate (SLS) positive control, 200 ug/mL | CNTLAGAMT | Control Agent amount (levels) | 110 | LSL-200 | sodium lauryl sulphate (SLS) positive control, 200 ug/mL | CNTLAGUNIT | Control Agent Unit | ug/mL | | Expand |
|---|
|
Row | STUDYID | DOMAIN | ENID (Entity ID) | ST vs. LT | Activation (+S9, -S9) | RUNID (Run-Port Number) | SMPID (Sample ID) | SMKFID (Smoke Fraction) | REPLCTID (Replicate Number) | PLATEID (Plate ID) | COLID (Column number) | ROWID (Row number) | SETCD (Set Code, TX) | RFSTDTC | RFENDTC | RFXSTDTC | RFXENDTC | RFCSTDTC | RFCENDTC | ARMCD | 1 | MA99999 | EI | 1 | 1-1 | 030001 | A | 1 | 1 | 1 | A | CS-10 | 2008-04-01T06:00 | | 2008-04-02T14:00 | 2008-04-02T14:04 | 2008-04-01T06:00 | 2008-04-01T06:00 | 3 | 2 | MA99999 | EI | 2 | 1-1 | 030001 | A | 1 | 1 | 1 | B | CS-10 | 2008-04-01T06:00 | 2008-04-21T14:00 | 2008-04-02T20:00 | 2008-04-02T20:04 | 2008-04-01T06:00 | 2008-04-01T06:00 | 3 |
| Dataset wrap |
|---|
|
| Rowcaps |
|---|
Rows 1-3, 8: | Show percentage result values that apply to GTREFID=C0. REFID=C0, as shown in the RELREF dataset, relates this data to the trial set in the first row of table 1 in the sample report table for study 123. | | Rows 4-7: | Show the 4 micronucleated cell counts for the observational units with GTREFID from C0-Count1 through C0-Count4, for which their relationship to test conditions (in tx.xpt) and experimental units (in relref.xpt) are shown in the RELREF dataset. | Rows 9-11, 16: | Show percentage result values that apply to GTREFID=C1250. REFID=C1250, as shown in the RELREF dataset, relates this data to the trial set in the second row of table 1 in the sample report table for study 123. | | Rows 12-15: | Show the 4 micronucleated cell counts for the observational units with GTREFID from C1250-Count1 through C1250-Count4, for which their relationship to test conditions (in tx.xpt) and experimental units (in relref.xpt) are shown in the RELREF dataset. |
|
| Dataset2 |
|---|
| Row | STUDYID | DOMAIN | GTSEQ | GTREFID | GTTESTCD | GTTEST | GTCELLEV | GTORRES | GTORRESU | GTCOLSRT | GTSTRESC | GTSTRESN | GTSTRESU | GTDTC |
|---|
| 1 | 123 | GT | 1 | C0 | | Relative Increase in Cell Count | 154 | 0 | % |
| 0 | 0 | % | 2022-05-25 | | 2 | 123 | GT | 2 | C0 | RCC | Relative Cell Count | 154 | 0 | % |
| 0 | 0 | % | 2022-05-25 | | 3 | 123 | GT | 3 | C0 | RPD | Relative Population Doubling | 154 | 0 | % |
| 0 | 0 | % | 2022-05-25 | | 4 | 123 | GT | 4 | C0-Count1 | MNCE | Micronucleated Cells | 2205 | 15 | |
| 15 | 15 |
| 2022-05-25 | | 5 | 123 | GT | 5 | C0-Count2 | MNCE | Micronucleated Cells | 2474 | 13 |
|
| 13 | 13 |
| 2022-05-25 | | 6 | 123 | GT | 6 | C0-Count3 | MNCE | Micronucleated Cells | 2758 | 17 |
|
| 17 | 17 |
| 2022-05-25 | | 7 | 123 | GT | 7 | C0-Count4 | MNCE | Micronucleated Cells | 2669 | 12 |
|
| 12 | 12 |
| 2022-05-25 | | 8 | 123 | GT | 8 | C0 | MNCECE | Micronucleated Cells/Total Cells |
| 0.57 | % |
| 0.57 | 0.57 | % | 2022-05-25 | | 9 | 123 | GT | 1 | C1250 | RICC | Relative Increase in Cell Count | 134 | 15.7 | % |
| 15.7 | 15.7 | % | 2022-05-25 | | 10 | 123 | GT | 2 | C1250 | RCC | Relative Cell Count | 134 | 13.0 | % |
| 13.0 | 13.0 | % | 2022-05-25 | | 11 | 123 | GT | 3 | C1250 | RPD | Relative Population Doubling | 134 | 7.9 | % |
| 7.9 | 7.9 | % | 2022-05-25 | | 12 | 123 | GT | 4 | C1250-Count1 | MNCE | Micronucleated Cells | 3266 | 20 |
|
| 20 | 20 |
| 2022-05-25 | | 13 | 123 | GT | 5 | C1250-Count2 | MNCE | Micronucleated Cells | 2190 | 17 |
|
| 17 | 17 |
| 2022-05-25 | | 14 | 123 | GT | 6 | C1250-Count3 | MNCE | Micronucleated Cells | 2758 | 13 |
|
| 13 | 13 |
| 2022-05-25 | | 15 | 123 | GT | 7 | C1250-Count4 | MNCE | Micronucleated Cells | 2714 | 21 |
|
| 21 | 21 |
| 2022-05-25 | | 16 | 123 | GT | 8 | C1250 | MNCECE | Micronucleated Cells/Total Cells |
| 0.66 | % |
| 0.66 | 0.66 | % | 2022-05-25 |
|
|
| Expand |
|---|
| title | Notes for creating GT domain |
|---|
|
- Entity ID links to EI domain (usubjid)
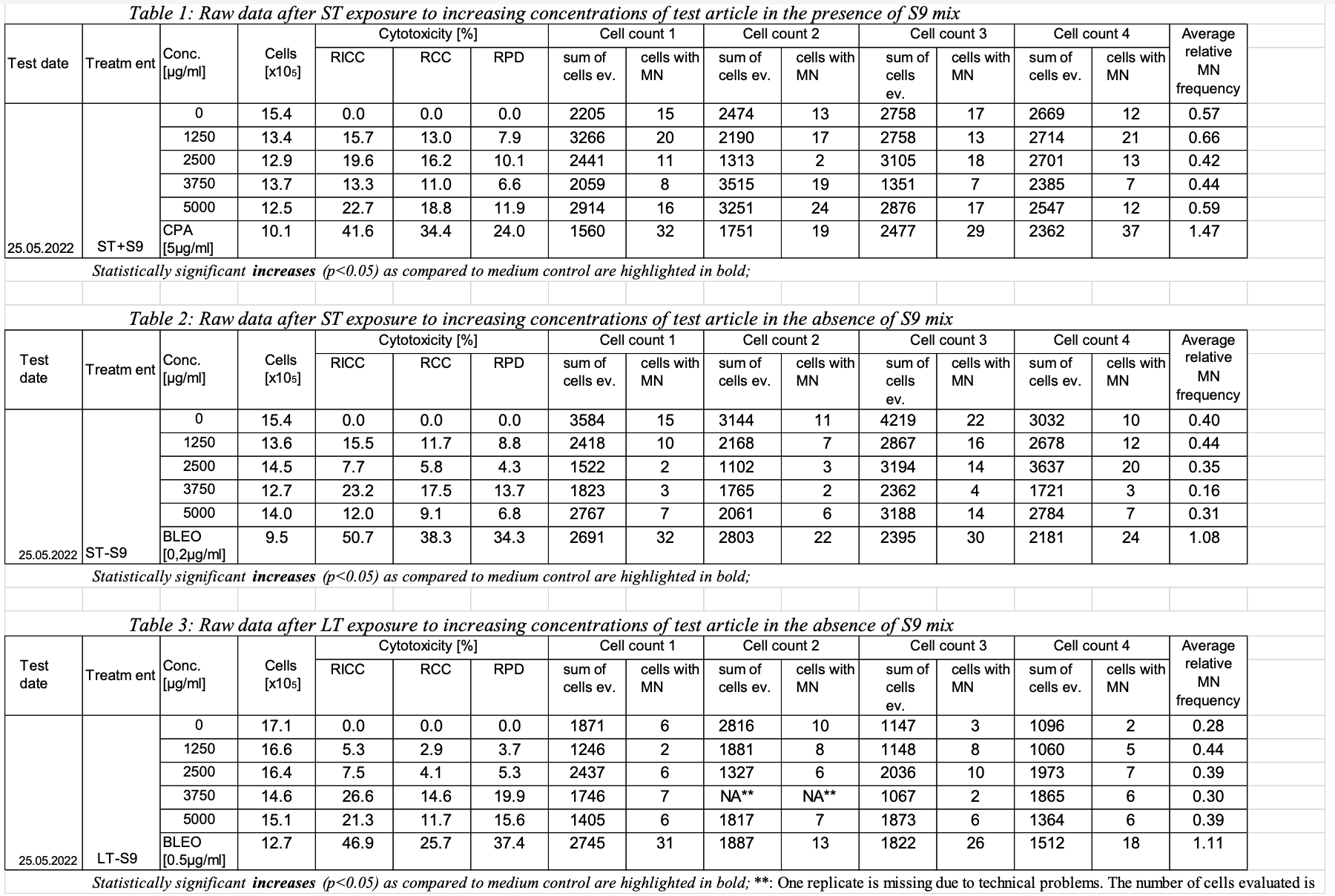
- TEST: After ST exposure to increasing concentrations of test article in the presence of S9 mix
| Expand |
|---|
| title | gt.xpt (similar to LB) |
|---|
|
|
| Row | STUDYID | DOMAIN | ENID (Entity ID) | GTSEQ | GTTESTCD | GTTEST | GTORRES | GTORRESU | GTSTRESC | GTSTRESN | GTSTRESU | GTSPEC??? | GTMETHOD | GTDTC | GTDY | GTNOMDY | GTELTM | GTTPTREF |
|---|
1 | ST487-12 | GT | 1 | MNT | In vitro Micronucleus Assay | ug/mL | ug/mL | 2 | ST487-12 | GT | 2 | MNT | In vitro Micronucleus Assay | ug/mL | ug/mL | 3 | ST487-12 | GT | 3 | MNT | In vitro Micronucleus Assay | ug/mL | ug/mL | 4 | ST487-12 | GT | 4 | MNT | In vitro Micronucleus Assay | ug/mL | ug/mL | 5 | ST487-12 | GT | 5 | MNT | In vitro Micronucleus Assay | ug/mL | ug/mL | 6 | ST487-12 | GT | 6 | MNT | In vitro Micronucleus Assay | ug/mL | ug/mL | 7 | ST487-12 | GT | 7 | MNT | In vitro Micronucleus Assay | ug/mL | ug/mL | 8 | ST487-12 | GT | 8 | MNT | In vitro Micronucleus Assay | ug/mL | ug/mL | 9 | ST487-12 | GT | 9 | MNT | In vitro Micronucleus Assay | ug/mL | ug/mL